CIESC Journal ›› 2020, Vol. 71 ›› Issue (2): 724-735.DOI: 10.11949/0438-1157.20190862
• Biochemical engineering and technology • Previous Articles Next Articles
Xue PENG( ),Chenlin LU,Diannan LU(
),Chenlin LU,Diannan LU( )
)
Received:2019-07-30
Revised:2019-09-03
Online:2020-02-05
Published:2020-02-05
Contact:
Diannan LU
通讯作者:
卢滇楠
作者简介:彭雪(1994—),女,博士研究生,基金资助:CLC Number:
Xue PENG, Chenlin LU, Diannan LU. Investigation on migration process of oxygen and carbon monoxide in human hemoglobin[J]. CIESC Journal, 2020, 71(2): 724-735.
彭雪, 芦琛璘, 卢滇楠. 氧气和一氧化碳在人血红蛋白迁移过程研究[J]. 化工学报, 2020, 71(2): 724-735.
Add to citation manager EndNote|Ris|BibTeX
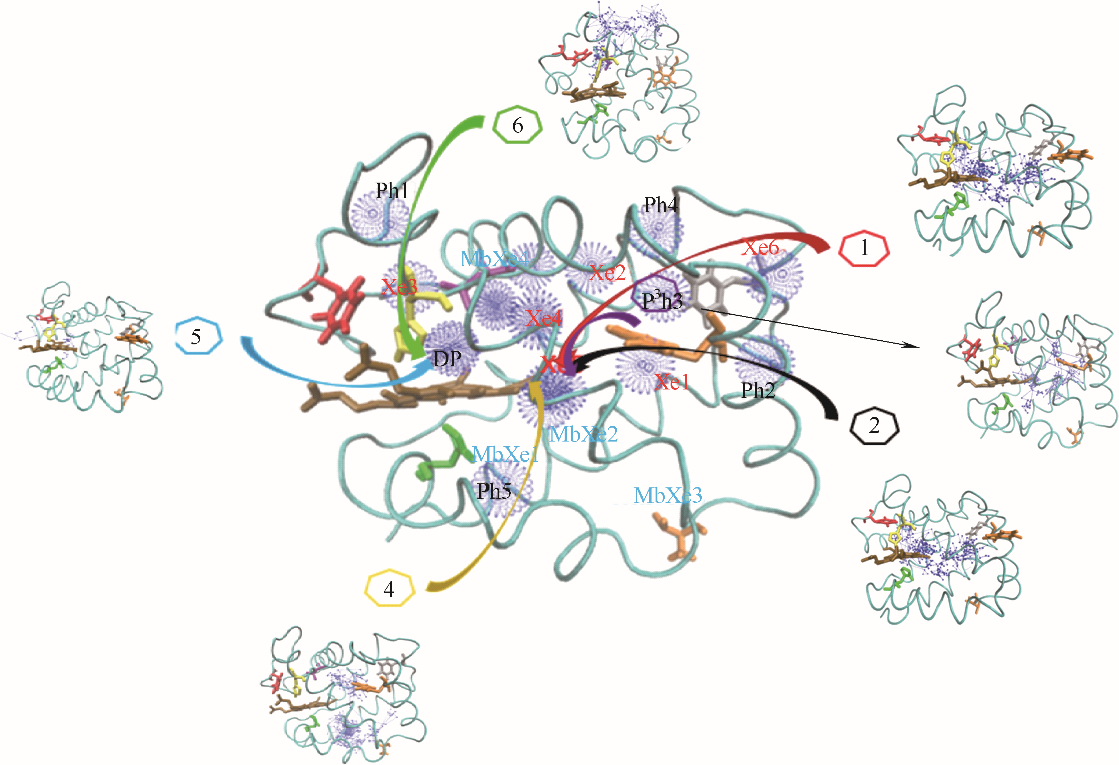
Fig.1 O2 binding sites and 6 major migration pathways in α chain of human hemoglobin(blue ball: O2 sites; line with arrow: migration trajectories of O2; cyan belt: backbone of α chain of human hemoglobin; brown stick: heme; Xe1—Xe6: Xe binding sites determined by experiments; DP: distal path site determined by experiemnts; Ph1— Ph5: instant cavities; MbXe1 — MbXe4: Xe binding sites of myoglobin determined by experiments)
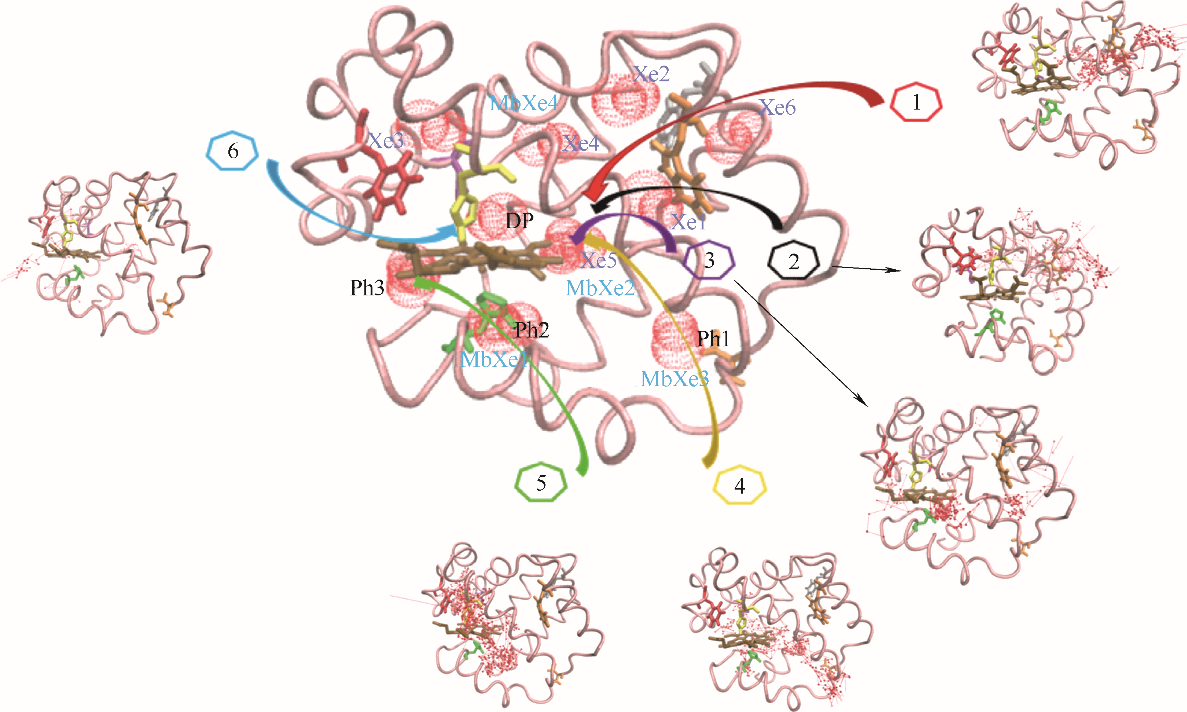
Fig.4 CO binding sites and 6 major migration pathways in β chain of human hemoglobin(red ball: CO sites; line with arrow: migration trajectories of CO; pink belt: backbone of β chain of human hemoglobin; brown stick: heme; Xe1—Xe6: Xe binding sites determined by experiments; DP: distal path site determined by experiemnts; Ph1—Ph3: instant cavities; MbXe1—MbXe4: Xe binding sites of myoglobin determined by experiments)
| 路径 | 次数 | 概率/% |
|---|---|---|
| 1: 水溶液→Xe6→Ph3→Xe5→DP | 49 | 49.0 |
| 2: 水溶液→Ph2→Xe1→Xe5→DP | 18 | 18.0 |
| 3: 水溶液→Ph3→Xe1→Xe5→DP | 3 | 3.0 |
| 4: 水溶液→Ph5→Xe5→Xe4→DP | 12 | 12.0 |
| 5: 水溶液→DP | 6 | 6.0 |
| 6: 水溶液→Ph1→Xe3→DP | 10 | 10.0 |
| 其他 | 2 | 2.0 |
Table 1 Frequency and probability of O2 migration pathways in α chain
| 路径 | 次数 | 概率/% |
|---|---|---|
| 1: 水溶液→Xe6→Ph3→Xe5→DP | 49 | 49.0 |
| 2: 水溶液→Ph2→Xe1→Xe5→DP | 18 | 18.0 |
| 3: 水溶液→Ph3→Xe1→Xe5→DP | 3 | 3.0 |
| 4: 水溶液→Ph5→Xe5→Xe4→DP | 12 | 12.0 |
| 5: 水溶液→DP | 6 | 6.0 |
| 6: 水溶液→Ph1→Xe3→DP | 10 | 10.0 |
| 其他 | 2 | 2.0 |
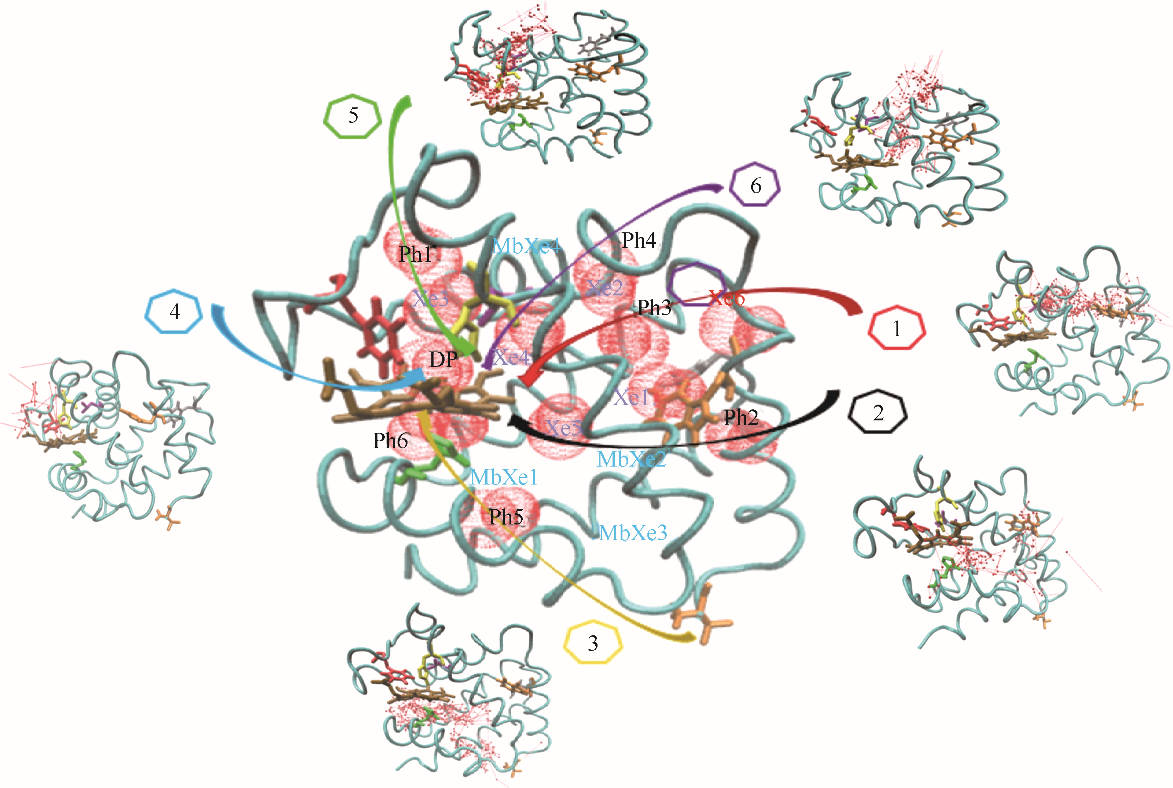
Fig.2 CO binding sites and 6 major migration pathways in α chain of human hemoglobin(red ball: CO sites; line with arrow: migration trajectories of CO; cyan belt: backbone of α chain of human hemoglobin; brown stick: heme; Xe1—Xe6: Xe binding sites determined by experiments; DP: distal path site determined by experiemnts; Ph1—Ph6: instant cavities; MbXe1—MbXe4: Xe binding sites of myoglobin determined by experiment)
| 路径 | 次数 | 概率/% |
|---|---|---|
| 1: 水溶液→Xe6→Ph3→Xe4→DP | 21 | 17.9 |
| 2: 水溶液→Xe6→Ph2→Xe5→DP | 5 | 4.3 |
| 3: 水溶液→Ph5→Ph6→DP | 5 | 4.3 |
| 4: 水溶液→DP | 34 | 29.0 |
| 5: 水溶液→Ph1→Xe3→DP | 37 | 31.6 |
| 6: 水溶液→Ph4→Xe2→Xe4→DP | 12 | 10.3 |
| 其他 | 3 | 2.6 |
Table 2 Frequency and probability of CO migration pathways in α chain
| 路径 | 次数 | 概率/% |
|---|---|---|
| 1: 水溶液→Xe6→Ph3→Xe4→DP | 21 | 17.9 |
| 2: 水溶液→Xe6→Ph2→Xe5→DP | 5 | 4.3 |
| 3: 水溶液→Ph5→Ph6→DP | 5 | 4.3 |
| 4: 水溶液→DP | 34 | 29.0 |
| 5: 水溶液→Ph1→Xe3→DP | 37 | 31.6 |
| 6: 水溶液→Ph4→Xe2→Xe4→DP | 12 | 10.3 |
| 其他 | 3 | 2.6 |

Fig.3 O2 binding sites and 6 major migration pathways in β chain of human hemoglobin(blue ball: O2 sites; line with arrow: migration trajectories of O2; pink belt: backbone of β chain of human hemoglobin; brown stick: heme; Xe1—Xe6: Xe binding sites determined by experiments; DP: distal path site determined by experiemnts; Ph1: instant cavities; MbXe1 — MbXe4: Xe binding sites of myoglobin determined by experiments)
| 路径 | 次数 | 概率/% |
|---|---|---|
| 1: 水溶液→Xe6→Xe1→Xe5→DP | 25 | 25.0 |
| 2: 水溶液→Xe5→DP | 11 | 11.0 |
| 3: 水溶液→Xe1→Xe5→DP | 36 | 36.0 |
| 4: 水溶液→Ph1→Xe5→→DP | 5 | 5.0 |
| 5: 水溶液→DP | 4 | 4.0 |
| 6: 水溶液→Xe2→Xe4→DP | 16 | 16.0 |
| 其他 | 3 | 3.0 |
Table 3 Frequency and probability of O2 migration pathways in β chain
| 路径 | 次数 | 概率/% |
|---|---|---|
| 1: 水溶液→Xe6→Xe1→Xe5→DP | 25 | 25.0 |
| 2: 水溶液→Xe5→DP | 11 | 11.0 |
| 3: 水溶液→Xe1→Xe5→DP | 36 | 36.0 |
| 4: 水溶液→Ph1→Xe5→→DP | 5 | 5.0 |
| 5: 水溶液→DP | 4 | 4.0 |
| 6: 水溶液→Xe2→Xe4→DP | 16 | 16.0 |
| 其他 | 3 | 3.0 |
| 路径 | 次数 | 概率/% |
|---|---|---|
| 1: 水溶液→Xe6→Xe1→Xe5→DP | 14 | 13.0 |
| 2: 水溶液→Xe1→Xe2→Xe4→DP | 18 | 16.7 |
| 3: 水溶液→Xe1→Xe5→DP | 7 | 6.5 |
| 4: 水溶液→Ph1→Xe5→DP | 18 | 16.7 |
| 5: 水溶液→Ph2→Ph3→DP | 7 | 6.4 |
| 6: 水溶液→DP | 40 | 37.0 |
| 其他 | 4 | 3.7 |
Table 4 Frequency and probability of CO migration pathways in β chain
| 路径 | 次数 | 概率/% |
|---|---|---|
| 1: 水溶液→Xe6→Xe1→Xe5→DP | 14 | 13.0 |
| 2: 水溶液→Xe1→Xe2→Xe4→DP | 18 | 16.7 |
| 3: 水溶液→Xe1→Xe5→DP | 7 | 6.5 |
| 4: 水溶液→Ph1→Xe5→DP | 18 | 16.7 |
| 5: 水溶液→Ph2→Ph3→DP | 7 | 6.4 |
| 6: 水溶液→DP | 40 | 37.0 |
| 其他 | 4 | 3.7 |
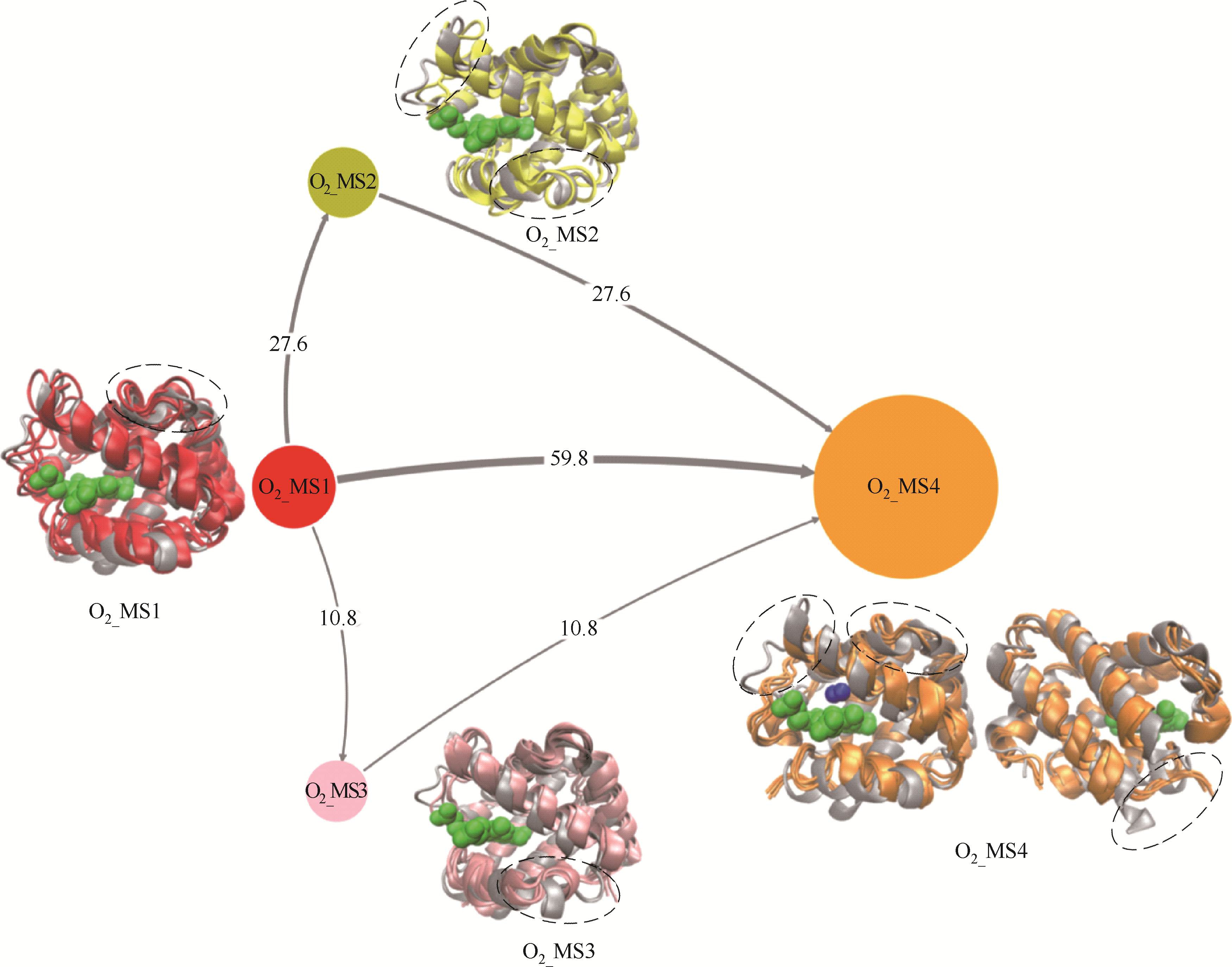
Fig.7 Tructural transition pathways of α chain of human hemoglobin during O2 migration processes(gray ribbon model: crystal structure of α chain of human hemoglobin; O2_MS1 in red: initial structure, i.e., O2 unbinding state; O2_MS2 in yellow and O2_MS3 in pink: middle states, i.e., key structures during O2 migration; O2_MS4 in orange: final O2 binding state; lines with arrow: transition pathways among different states, their thickness reflects transition probability; circles with different color: different states, their sizes reflect appearance probability; ribbon model with different color: corresponding structures of α chain of human hemoglobin; green ball model: heme molecule)
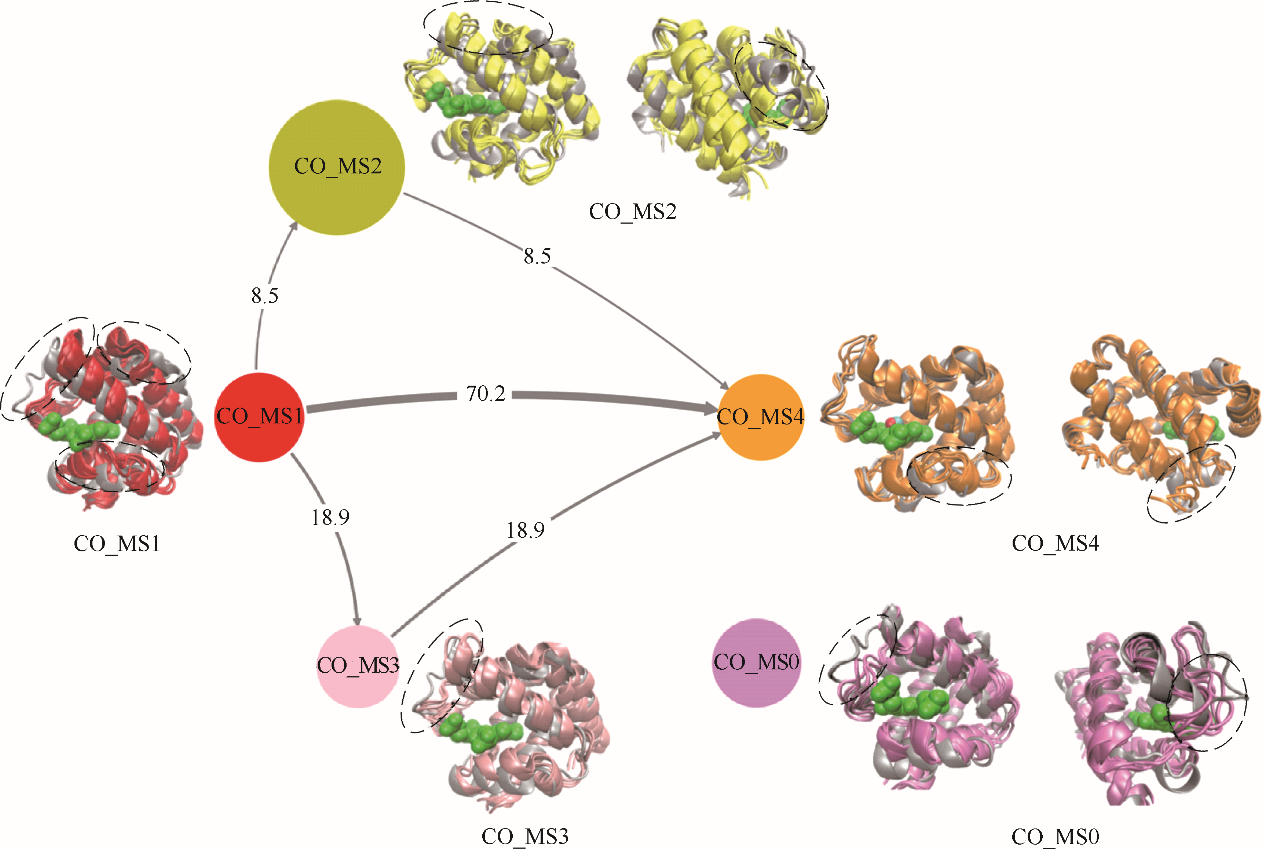
Fig.8 Structural transition pathways of α chain of human hemoglobin during CO migration processes(gray ribbon model: crystal structure of α chain of human hemoglobin; CO _MS1 in red: initial structure, i.e., CO unbinding state; CO _MS2 in yellow and CO _MS3 in pink: middle states, i.e., key structures during CO migration; CO _MS0 in purple: independent state, i.e., a structure not in transition network; CO _MS4 in orange: final CO binding state; lines with arrow: transition pathways among different states, their thickness reflects transition probability; circles with different color: different states, their sizes reflect appearance probability; ribbon model with different color: corresponding structures of α chain of human hemoglobin; green ball model: heme molecule)
| 1 | Feitelson J, McLendon G. Migration of small molecules through the structure of hemoglobin: evidence for gating in a protein electrontransfer reaction [J]. Bioc., 1991, 30: 5051-5055. |
| 2 | Leitner D M. Energy flow in proteins [J]. Annu. Rev. Phys. Chem., 2008, 59: 233-259. |
| 3 | Savino C, Miele A E, Draghi F, et al. Pattern of cavities in globins: the case of human hemoglobin [J]. Biopolymers, 2009, 91: 1097-1107. |
| 4 | Park S Y, Yokoyama T, Shibayama N, et al. 1.25 Å resolution crystal structures of human haemoglobin inthe oxy, deoxy and carbonmonoxy forms [J]. J. Mol. Biol., 2006, 360: 690-701. |
| 5 | Schotte F, Lim M, Jackson T, et al. Watching a protein as it functions with 150-ps time-resolved X-ray crystallography [J]. Science, 2003, 300: 1944-1947. |
| 6 | Schotte F, Soman J, Olson J S, et al. Picosecond time-resolved X-ray crystallography: probing protein function in real time [J]. Struct. Biol., 2004, 147: 235-246. |
| 7 | Schmidt M, Nienhaus K, Pahl R, et al. Ligand migration pathway and protein dynamics in myoglobin: a time-resolved crystallographic study on L29W MbCO [J]. Proc. Natl. Acad., 2005, 102: 11704-11709. |
| 8 | Perutz M F, Mathews F S. An X-ray study of azide methaemoglobin [J]. Biol., 1966, 21: 199- 202. |
| 9 | Lucas M F, Guallar V. An atomistic view on human hemoglobin carbon monoxide migration processes [J]. Biophys., 2012, 102: 887-896. |
| 10 | Lepeshkevich S V, Biziuk S A, Lemeza A M. The kinetics of molecular oxygen migration in the isolated alpha chains of human hemoglobin as revealed by molecular dynamics simulations and laser kinetic spectroscopy [J]. Biochimica et Biophysica Acta, 2011, 1814: 1279-1288. |
| 11 | Lepeshkevich S V, Gilevich S N, Parkhats M V, et al. Molecular oxygen migration through the xenon docking sites of human hemoglobin in the R-state [J]. Biochimica et Biophysica Acta (BBA) - Proteins and Proteomics, 2016, 1864(9): 1110-1121. |
| 12 | Hummer G, Schotte F, Anfinrud P A, et al. Unveiling functional protein motions with picosecond X-ray crystallography and molecular dynamics simulations [J]. Proceedings of the National Academy of Sciences of the United States of America, 2004, 101(43): 15330-15334. |
| 13 | Ruscio J Z, Kumar D, Shukla M. Atomic level computational identification of ligand migration pathways between solvent and binding site in myoglobin [J]. Proc. Natl., 2008, 105: 9204-9209. |
| 14 | Elber R. Ligand diffusion in globins simulations versu experiment [J]. Biol., 2010, 20: 162-167. |
| 15 | Ostermann A, Waschipky R, Parak F G, et al. Ligand binding and conformational motions in myoglobin [J]. Nature, 2000, 404(6774): 205-208. |
| 16 | Cordone L, Cottone G, Giuffrida S, et al. Thermal evolution of the CO stretching band in carboxy-myoglobin in the light of neutron scattering and molecular dynamics simulations [J]. Chemical Physics, 2008, 345(2): 275-282. |
| 17 | Yu T Q, Lapelosa M, Vandeneijnden E, et al. Full kinetics of CO entry, internal diffusion, and exit in myoglobin from transition-path theory simulations [J]. Journal of the American Chemical Society, 2015, 137(8): 3041-3050. |
| 18 | Alessandro B, Massimiliano B, Michele P. Wiley interdisciplinary reviews: computational molecular [J]. Journal of Computational Chemistry, 2011, 8: 826-843. |
| 19 | Leitner D M. Energy flow in proteins [J]. Annu. Rev. Phys. Chem., 2008, 59: 233-259. |
| 20 | Elber R, Karplus M. Enhanced sampling in molecular dynamics: use of the time-dependent hartree approximation for a simulation of carbon monoxide diffusion through myoglobin [J]. Journal of the American Chemical Society, 1990, 112: 9161-9175. |
| 21 | Meller J, Elber R. Computer simulations of carbon monoxide photodissociation in myoglobin: structural interpretation of the Bstates [J]. Biophys., 1998, 74: 789-802. |
| 22 | Bossa C, Anselmi M, Roccatano D, et al. Extended molecular dynamics simulation of thecarbon monoxide migration in sperm whale myoglobin [J]. Biophys., 2004, 86: 3855-3862. |
| 23 | Cohen J, Arkhipov A, Braun R, et al. Imaging the migration pathways for O2, CO, NO, and Xe inside myoglobin [J]. Biophysical Journal, 2006, 91(5): 1844-1857. |
| 24 | Sancho D D, Kubas A, Wang P H, et al. Identification of mutational hot spots for substrate diffusion: application to myoglobin [J]. Journal of Chemical Theory & Computation, 2015, 11(4): 1919-1927. |
| 25 | Shadrina M S, English A M, Peslherbe G H. Effective simulations of gas diffusion through kinetically accessible tunnels in multisubunit proteins: O2 pathways and escape routes in T-state deoxyhemoglobin [J]. J. Am. Chem. Soc., 2012, 134: 11177-11184. |
| 26 | Takayanagi M, Kurisaki I, Nagaoka M. Oxygen entry through multiple pathways in T-state human hemoglobin [J]. J. Phys. Chem. B, 2013, 117: 6082-6091. |
| 27 | Wang P H, Best R B, Blumberger J. Multiscale simulation reveals multiple pathways for H2 and O2 transport in a [NiFe]-hydrogenase [J]. Journal of the American Chemical Society, 2013, 10: 3548-3556. |
| 28 | Bustamante J P, Szretter M E, Sued M, et al. A quantitative model for oxygen uptake and release in a family of hemeproteins [J]. Bioinformatics, 2016, 32(12): 1805-1813. |
| 29 | Liang J, Edelsbrunner H, Fu P, et al. Analytical shape computation of macromolecules (Ⅱ): Inacessiblecavities in proteins [J]. Proteins: Struct. Func. Gen., 1998, 33: 18-29. |
| 30 | Shaanan B. Structure of human oxyhaemoglobin at 2.1 Å resolution [J]. Mol. Biol., 1983, 171: 31-59. |
| 31 | Wang P H, De S D, Best R B. Computation of rate constants for diffusion of small ligands to and from buried protein active sites [J]. Methods in Enzymology, 2016, 578: 299-326. |
| 32 | Birukou I, Maillett D H, Birukova A. Modulating distal cavities in the α and β subunits of human HbA reveals the primary ligand migration pathway [J]. Biochemistry, 2011, 50(34): 7361-7374. |
| 33 | Takayanagi M, Nagaoka M. Incipient structural and vibrational relaxation process of photolyzed carbonmonoxy myoglobin: statistical analysis by perturbation ensemble molecular dynamics method [J]. Theor. Chem. Acc., 2011, 130: 1115-1129. |
| 34 | Estarellas M C, Seira C C, Luque F J, et al. Understanding the kinetics of ligand binding to globins with molecular dynamics simulations: the necessity of multiple state models [J]. Drug Discovery Today Technologies, 2015, 17(70): 22-27. |
| 35 | Brooks B R, Bruccoleri R E, Olafson B D, et al. CHARMM: a program for macromolecular energy, minimization, and dynamics calculations [J]. Comp. Chem., 1983, 4: 187-217. |
| 36 | Shadrina M S, Peslherbe G H, English A M. O2 and water migration pathways between the solvent and heme pockets of hemoglobin with open and closed conformations of the distal hisE7 [J]. Biochemistry, 2015, 54(34): 5279-5289. |
| 37 | Leitner D M, Straub J E. Proteins: Energy, Heat and Signal Flow (Computation in Chemistry) [M]. Boca Raton, FL: CRC Press, 2009. |
| 38 | Cameron A, Giovanni B. Enhanced sampling in molecular dynamics using metadynamics, replica exchange, and temperature-acceleration [J]. Entropy, 2013, 16: 163-199. |
| 39 | Berenbrink M. Evolution of vertebrate haemoglobins: histidine side chains, specific buffer value and Bohr effect [J]. Respiratory Physiology & Neurobiology, 2006, 154(1): 165-184. |
| 40 | Huber K P, Herzberg G. Molecular Spectra and Molecular Structure [M]. US: Springer, 1979. |
| [1] | Mingkun XIAO, Guang YANG, Yonghua HUANG, Jingyi WU. Numerical study on bubble dynamics of liquid oxygen at a submerged orifice [J]. CIESC Journal, 2023, 74(S1): 87-95. |
| [2] | Minghao SONG, Fei ZHAO, Shuqing LIU, Guoxuan LI, Sheng YANG, Zhigang LEI. Multi-scale simulation and study of volatile phenols removal from simulated oil by ionic liquids [J]. CIESC Journal, 2023, 74(9): 3654-3664. |
| [3] | Jianbo HU, Hongchao LIU, Qi HU, Meiying HUANG, Xianyu SONG, Shuangliang ZHAO. Molecular dynamics simulation insight into translocation behavior of organic cage across the cellular membrane [J]. CIESC Journal, 2023, 74(9): 3756-3765. |
| [4] | Jiajia ZHAO, Shixiang TIAN, Peng LI, Honggao XIE. Microscopic mechanism of SiO2-H2O nanofluids to enhance the wettability of coal dust [J]. CIESC Journal, 2023, 74(9): 3931-3945. |
| [5] | Shaoqi YANG, Shuheng ZHAO, Lungang CHEN, Chenguang WANG, Jianjun HU, Qing ZHOU, Longlong MA. Hydrodeoxygenation of lignin-derived compounds to alkanes in Raney Ni-protic ionic liquid system [J]. CIESC Journal, 2023, 74(9): 3697-3707. |
| [6] | Linzheng WANG, Yubing LU, Ruizhi ZHANG, Yonghao LUO. Analysis on thermal oxidation characteristics of VOCs based on molecular dynamics simulation [J]. CIESC Journal, 2023, 74(8): 3242-3255. |
| [7] | Feifei YANG, Shixi ZHAO, Wei ZHOU, Zhonghai NI. Sn doped In2O3 catalyst for selective hydrogenation of CO2 to methanol [J]. CIESC Journal, 2023, 74(8): 3366-3374. |
| [8] | Ming DONG, Jinliang XU, Guanglin LIU. Molecular dynamics study on heterogeneous characteristics of supercritical water [J]. CIESC Journal, 2023, 74(7): 2836-2847. |
| [9] | Ji CHEN, Ze HONG, Zhao LEI, Qiang LING, Zhigang ZHAO, Chenhui PENG, Ping CUI. Study on coke dissolution loss reaction and its mechanism based on molecular dynamics simulations [J]. CIESC Journal, 2023, 74(7): 2935-2946. |
| [10] | Yuanchao LIU, Xuhao JIANG, Ke SHAO, Yifan XU, Jianbin ZHONG, Zhuan LI. Influence of geometrical dimensions and defects on the thermal transport properties of graphyne nanoribbons [J]. CIESC Journal, 2023, 74(6): 2708-2716. |
| [11] | Xiaowen ZHOU, Jie DU, Zhanguo ZHANG, Guangwen XU. Study on the methane-pulsing reduction characteristics of Fe2O3-Al2O3 oxygen carrier [J]. CIESC Journal, 2023, 74(6): 2611-2623. |
| [12] | Ruikang LI, Yingying HE, Weipeng LU, Yuanyuan WANG, Haodong DING, Yongming LUO. Study on the electrochemical enhanced cobalt-based cathode to activate peroxymonosulfate [J]. CIESC Journal, 2023, 74(5): 2207-2216. |
| [13] | Hao GU, Fujian ZHANG, Zhen LIU, Wenxuan ZHOU, Peng ZHANG, Zhongqiang ZHANG. Desalination performance and mechanism of porous graphene membrane in temporal dimension under mechanical-electrical coupling [J]. CIESC Journal, 2023, 74(5): 2067-2074. |
| [14] | Chenxin LI, Yanqiu PAN, Liu HE, Yabin NIU, Lu YU. Carbon membrane model based on carbon microcrystal structure and its gas separation simulation [J]. CIESC Journal, 2023, 74(5): 2057-2066. |
| [15] | Jiajing BAO, Hongfei BIE, Ziwei WANG, Rui XIAO, Dong LIU, Shiliang WU. The effects of adding long-chain ethers in n-heptane counterflow diffusion flames on the formation characteristics of soot precursors [J]. CIESC Journal, 2023, 74(4): 1680-1692. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||